|
I was notified this morning by Dr. Tracy Heath, Awards Director of the Society of Systematic Biologists (SSB), that I am awarded a Mini-ARTS grant ($1000). This award will allow me to travel to the Natural History Museum (NHM), UK to examine, digitize and document type specimens of species of the "Exophthalmus genus complex" (EGC). These are broad-nosed weevils (Curculionidae: Entiminae) from primarily Central America and the West Indies. The museum visit is needed for my project on the reclassification of the EGC. My collaborator and mentor at NHM is Dr. Chris Lyal, who is a world-renowned weevil taxonomist and co-authored the catalogue of genus and family group names of Curculionoidea. According to the description on SSB's website, "the Mini-ARTS grants are for revisionary taxonomy and systematics, modeled after the NSF Dear Colleague Letter: Advancing Revisionary Taxonomy and Systematics (ARTS) recently developed within the Systematics and Biodiversity Science Cluster." Their intended use is to allow SSB members to "spend a summer or semester apprenticed to an expert in a particular taxonomic group or to enhance revisionary taxonomic and systematics research in novel ways." I am very excited about this award and much thankful to SSB for setting funds to make this award. The Mini-ARTS grant is a positive signal from an important society, the SSB, that revisionary taxonomy and fundamental classification work are still very much needed in the postgenomics era. Among comparative biologists, there is little doubt that phylogenetic trees are indispensable powerful popular instruments for comparative biological investigations and it is also undeniable that computational phylogenetic research has risen to prominence within SSB. However, we are repeatedly realizing and re-realizing that species identities and the expression and communication of phylogenetic relationships in the form of a classification will continue to be a central theme of the systematic biology of all organisms. My award proposal is available for viewing and downloading
0 Comments
I have a morphological character matrix previously used in a cladistic analysis. I now would like to use this data set to generate taxonomic descriptions and identification keys. It turned out this is not really a straightforward process. The matrix was stored in Winclada. I was able to export a nexus file of the data set. I would to use DELTA to edit this data set. DELTA does not read a nexus file. After some research I was able to import the nexus file into DELTA. See the procedure described below.
Download NDE (Nexus Data Editor) here (runs on Windows 7, but did not run on my Windows 8 desktop) written by Rod Page. Open nexus file in NDE and go to 'Export' (under 'File'). Select 'DELTA' under Export. A pop-up window appears. Select directory/folder to export the DELTA format 'directive files' (there are four of them). Put these files in a new folder. In DELTA select 'Import directives...' (under 'File'). Copy/paste the folder directory (folder path) where you just saved the DELTA format directive files. A new window pops up and click 'OK'. Your data set should now appear in DELTA Editor. Also see discussions at the DELTA mail-list about format conversion: [Converting Nexus to DELTA] [Nexus vs DELTA files] [Delta, Lucid and the Nexus data editor] [Nexus data editor for Windows] [add different sets of attributes to items] I advised three undergraduate research students past semester. They were involved in aspects of my research project on the phylogenetics and biogeography of a group of weevils. The students were assigned individual projects focusing a particular task/aspect. One student performed molecular work, doing lots of PCR and DNA sequencing. Another student was mainly databasing and imaging weevil specimens. The last student worked on weevil specimens preserved in alcohol, having to sort them to species.
To learn about their experiences of doing research in the lab and with me, I invited the students for a casual gathering today. And there are a few things for future consideration. 1. The students would like to have some general ideas on how their individual projects fit to the bigger of picture of research. This was brought up because the individual projects only concerned very specific aspects of our research program and it was hard for a student to see why he/she was doing a particular task, e.g., PCR or specimen-sorting. Although I had always described the bigger project at the beginning, I guess I did not realize it was difficult for the students to understand the relevance of his/her project just by listening to a short verbal description. Two things could be done in the future. First, make the description more concrete. Provide more examples. Second, reiterate the goal(s) of the project and re-explain the project several times as the student proceeds with his/her research. Students gain greater and better understanding of their research/work after some hands-on experience, and iterations would help them further their understands of their positions in the research program. 2. The students would like to have some general lectures on the research program conducted in the lab. Our lab works on the taxonomy, systematics, phylogenetics and evolution of weevils. Students would benefit from learning the basics of these subjects. Taxonomy is not a well-covered subject in university biology/life sciences curriculum nowadays. Some of the students did not have a good understanding of Linnaean taxonomy. It turned out to be quite hard to communicate with students about the concepts of taxonomic ranks such as order, family, subfamily and genus, which are essential for virtually any aspects of our research program. Perhaps learning the basic concepts would help. 3. One student mentioned he would like to have more technical help//guidance on a particular task. I have to admit that there was some lack of supervision from my side in this particular student. Periodical discussions on research or technical challenges would resolve this issue in the future. 4. One student commented that he would like to do a greater variety of work. He was mainly doing molecular work, which happens to be rather standardized. This is somewhat difficult to address because it would be hard to have a novice undergraduate student multi-task at the beginning of his/her research. Other means/methods are needed to sustain a student's interested and motivation. See No. 5 below. 5. Concrete products/results may help students sustain their interest in the work. The same student who raised the issue described in No.4 also commented that seeing actual gel pictures (for visualizing PCR results) was always something exciting and editing the chromatograms (of DNA sequences) was also something enjoyable. These are visually concrete products that a person can interact with directly (as compared to the invisible DNA). The other two students did not have this problem because they worked directly with actual weevil specimens. 6. In relation to No. 5, it would be good to lay out clear project/research goals and have appropriate assessment methods. It is easy to get disinterested or simply habituated to keep doing a certain task without really knowing what one has achieved. For future students, I will try setting goals for them in terms of the skills to gain, the tasks to accomplish and the knowledge to learn, and assess their research/learning progress frequently. 7. Assess strengths and weaknesses of students and assign projects accordingly. This requires some experience working together with students and making observations. But these may not be enough. During our meeting today, it turned out one student actually has an interest in photography and I did not know about it (even after working with him for a semester). I had him sort specimens most of the time. In future I can try doing some diagnostics on the student before assigning work. Overall, the students enjoyed their experiences doing research in the lab. It was also a great experience for me. Dealing with the diversity of and the differences among the students is probably the most challenging aspect of my mentoring. Still a lot more to learn and experiment with for myself. I will be attending and presenting at the ESA (Entomological Society of America) meeting in Austin, Texas the following week (Nov09-13). I will be giving two talks, one of the “Featured Young Professional Presentation” at the SysEB Section Meeting and another on my recent work on Caribbean weevils and biogeography. I will be also co-moderating a session on Wed morning (Nov 13) entitled "Systematics of Coleoptera and Biodiversity Technology".
Check out the cool poster below specially made for this year's meeting. Nice integration of science and arts. Phylogenetic analysis is a very computing-intensive process and very often requires supercomputing facilities. The CIPRES (Cyberinfrastructure for Phylogenetic Research) supercomputing facility based in San Diego is one of the most powerful facilities to run phylogenetic analyses for large data sets, and it is FREE! A variety of programs are available. They generally work quite well. However, one the programs frequently used for molecular dating, called 'BEAST', runs rater slow at this site. A data set of 130 taxa running for 10-million generations would take 7 hours to finish. This is even longer than running on a powerful desktop computer such as a PowerMac or a PC tower. What's more, a BEAST analysis are usually run for a 100 million generations, which would require 70 hours or four days for the CIPRES site to complete. Multiple runs are generally needed to adjust input parameters and at this speed a thorough exploration of one's data set would require up to a month of a few months.
Alternatives would be setting up access to a local cluster service. Many universities have such facilities, but often charge a fee to use. Last year I wrote a blog about manuscript review status. Here I report manuscript review status updates from another journal. Interestingly, the status updates are somewhat different from the ones that I wrote about in the previous blog. It seems every journal has slightly different systems.
1. Awaiting admin processing - This is the status I saw immediately after submission (Feb 23, 2013). the manuscript is being considered for the peer-review process. After submission the manuscript usually goes to the Editor in Chief or managing editor first. This person may or may not select reviewers for the manuscript. If there are subject editors or Associate Editors (AE), the manuscript may get assigned to one of them by the Editor in Chief or managing editor. The AE will then be in charge of handling the manuscript (inviting reviewers, collecting reviews and making acceptance/rejection recommendations). 2. Awaiting Referee Selection - The editor (an Associate Editor or the Editor in Chief) is in the process of inviting reviewers. However, at this stage, the manuscript may still get desk rejected. About two days after I submitted my manuscript, the status has changed to 'awaiting referee selection'. 3. Awaiting Referee Assignment - The editor has sent invitations to reviewers (referees) and is waiting for their responses to accept or decline the invitation. I saw this status on Mar 03, 2013, about 10 days after the submission. 4. Awaiting Referee Scores - Reviewers have started reviewing the manuscript. Th review time varies from journal to journal, with many having a one-month period. I saw this status on Mar 05, 2013, only two days after the 'awaiting referee assignment' status came up. 5. On March 16, or about 11 days after the reviewer started doing the reviews, I received an email from the editor, accepting my manuscript. On ManuscriptCentral, it was showing: Minor Revision, with Apr-16-2013 as the due date. I also found a short, but useful explanation of manuscript review status from the journal 'Molecular Medicine'. Awaiting Admin Processing – the manuscript is being considered for the peer-review process. Under Consideration – the manuscript has been sent for peer-review. Awaiting AE Preliminary Decision – peer-review reports have been received. Awaiting EDB Decision Approval – the manuscript is in the queue for Editorial Board Review and decision. You can also look up another useful blog (in Chinese, translate it using Google Translate). As a scientist, I have to read a lot to keep up with the literature and to acquire knowledge. Sometimes I find it frustrating how easily I forget what I read. I decided to keep simple notes of the readings I did and publish them on my blog so that they could be shared with other people. Here is the first in the reading notes series.
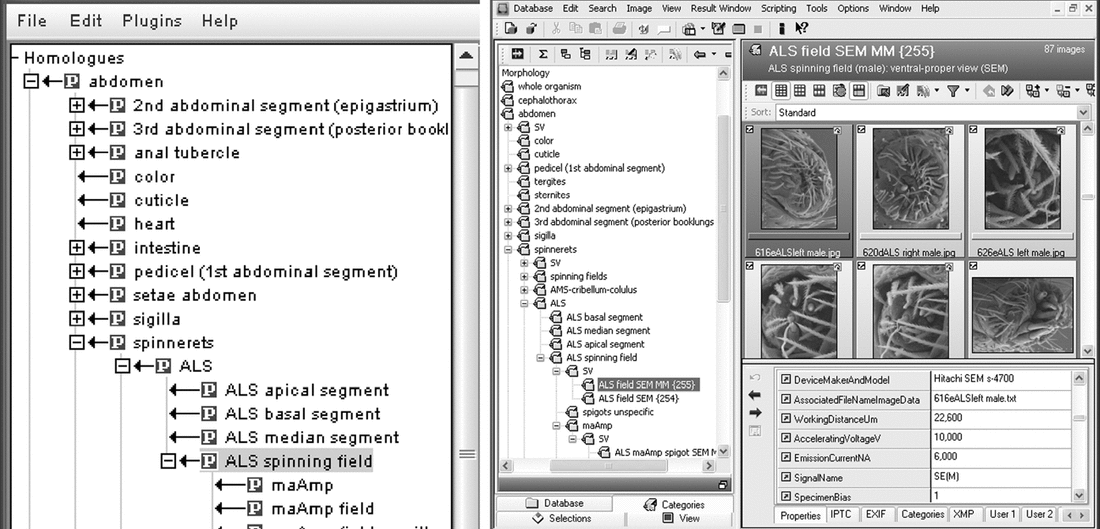
The article: Ramírez, M.J., Coddington, J.A., Maddison, W.P., Midford, P.E., Prendini, L., Miller, J., Griswold, C.E., Hormiga, G., Sierwald, P., Scharff, N., et al. (2007). Linking of digital images to phylogenetic data matrices using a morphological ontology. Syst Biol 56, 283–294. Abstract: Images are paramount in documentation of morphological data. Production and reproduction costs have traditionally limited how many illustrations taxonomy could afford to publish, and much comparative knowledge continues to be lost as generations turn over. Now digital images are cheaply produced and easily disseminated electronically but pose problems in maintenance, curation, sharing, and use, particularly in long-term data sets involving multiple collaborators and institutions. We propose an efficient linkage of images to phylogenetic data sets via an ontology of morphological terms; an underlying, fine-grained database of specimens, images, and associated metadata; fixation of the meaning of morphological terms (homolog names) by ostensive references to particular taxa; and formalization of images as standard views. The ontology provides the intellectual structure and fundamental design of the relationships and enables intelligent queries to populate phylogenetic data sets with images. The database itself documents primary morphological observations, their vouchers, and associated metadata, rather than the conventional data set cell, and thereby facilitates data maintenance despite character redefinition or specimen reidentification. It minimizes reexamination of specimens, loss of information or data quality, and echoes the data models of web-based repositories for images, specimens, and taxonomic names. Confusion and ambiguity in the meanings of technical morphological terms are reduced by ostensive definitions pointing to features in particular taxa, which may serve as reference for globally unique identifiers of characters. Finally, the concept of standard views (an image illustrating one or more homologs in a specific sex and life stage, in a specific orientation, using a specific device and preparation technique) enables efficient, dynamic linkage of images to the data set and automatic population of matrix cells with images independently of scoring decisions. Main ideas/results: 1. The authors identifies current problems with documenting and communicating morphological characters. 2. They propose a system and present a software (SILK) to "addresses the challenge of how to maintain, curate, share, and make efficient use of these collections of digital images". 3. They advocate the typification of morphological characters. Quotes: Note: Words or phrases in square brackets are my notes to the quoted sentences. Also, the quotes are taken at various parts of the article and do not show any connections among them. [Proposal of paper] We propose an efficient linkage of images to phylogenetic data sets via an ontology of morphological terms; an underlying, fine-grained database of specimens, images, and associated metadata; fixation of the meaning of morphological terms (homolog names) by ostensive references to particular taxa; and formalization of images as standard views. [Utilities/features] [1] The ontology provides the intellectual structure and fundamental design of the relationships and enables intelligent queries to populate phylogenetic data sets with images. [2] The database itself documents primary morphological observations, their vouchers, and associated metadata, rather than the conventional data set cell, and thereby facilitates data maintenance despite character redefinition or specimen reidentification. [3] It minimizes reexamination of specimens, loss of information or data quality, and echoes the data models of web-based repositories for images, specimens, and taxonomic names. [4] Confusion and ambiguity in the meanings of technical morphological terms are reduced by ostensive definitions pointing to features in particular taxa, which may serve as reference for globally unique identifiers of characters. [Current problems] Inability to document and disseminate morphological data, in turn, led to huge losses of comparative knowledge as generations turned over. Successive generations of specialists had to reevaluate that information. Information could not be efficiently preserved or disseminated. [Desirable properties of an image repository] [T]he repository should illustrate and justify the actual scores in matrix cells, as well as the concepts underlying each character and its states. It should offer queries of images based on homology hypotheses, and, ideally, facilitate discovery of more refined homology hypotheses. It should also be designed for collaboration and parallel workflows, integrating work of individual researchers and research groups into a common, publicly available repository that links images to phylogenetic studies. [My summary of the desirable properties of the image repository]
[Objective of paper] This paper addresses the challenge of how to maintain, curate, share, and make efficient use of these collections of digital images. BOTTLENECKS IN THE DOCUMENTATION, REPLICABILITY AND ACCUMULATION OF MORPHOLOGICAL DAT [five points as follows]
[Some problems of current practices in scoring and documenting morphological characters]
Ideally, every cell in a morphological data matrix should derive from an investigator-credited observation, and nearly all should be photo-documented in order to minimize the chance that future workers will need to repeat the observation and to maximize longevity and value of the data. [Character scoring process] [Two main methods:]
A "transverse" pass (scoring one character for all terminals), on the other hand, requires the preparation and manipulation of many specimens and is in general inefficient. Many experimental characters will be discarded or redefined as the study progresses, requiring multiple transverse passes. Storing and retrieving primary observations, especially images, can make transverse passes faster, because specimens are handled only once. [Character image acquisition for all cells] Attaching images to cells before scoring is not a trivial task. The Spider AToL project envisages a data set of 500 terminals by 1000 to 2000 characters, implying 500,000 to 1,000,000 cells, most of which ideally would be photodocumented (thus megabytes of data per cell). [Utility of images] Formulation of new character hypotheses requires examination of relevant images across many terminals. [Characters without typification] However, names of homologues (characters and character states) are not currently "fixed" by any sort of typification procedure, and, not coincidentally, their meanings are subject to eternal debate. [What is holotype] A holotype does not "define" a species scientifically, it merely provides the objective reference for a name to enable accurate communication. [Character typification- what not to do] [I]t would be unwise to fix homologue definitions by literally designating particular specimens as types or to ape the rules of taxonomic nomenclature. Links: http://research.amnh.org/atol/files/ Figures: The authors have propose a scheme to link multiple databases. I have not seen anything like what they depict in the figure. Perhaps something to consider? Some provoking thoughts on scientific literature.
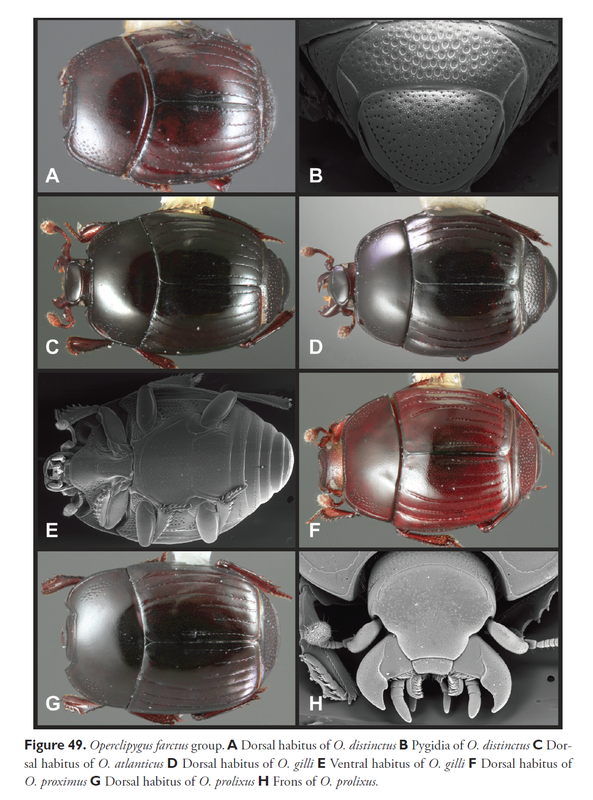
Also see a blog post from the 'ispider' site: Science is a Product in the Wrong Marketplace http://ispiders.blogspot.com/2012/01/science-is-product-in-wrong-marketplace.html Two entomologists at the Santa Barbara Museum of Natural History, Michael Caterino and Alexey Tishechkin have just described 138 new species of beetles in a single publication.
The article is entitled "A systematic revision of Operclipygus Marseul (Coleoptera, Histeridae, Exosternini" and is published in the journal Zookeys, a premier open access online journal with a focus on zoological systematics. Histeridae are rather small, usually a few millimeters in size. They are commonly called clown beetles or hister beetles. To me, their appearances are quite absurd. The lab led by Dr. Norman Johnson at the Ohio State University has just published their research website. I find it elegant, informative and quite pleasant to visit. There are pictures and video of field trips, PDF copies of publications, descriptions of research projects, information of researchers and useful links to other entomology or taxonomy related sites. It seems still under construction and I look forward to seeing a more complete version.
URL of the Johnson Lab website: http://osuc.osu.edu/Publish/index.html# Or click HERE [Correction: in the blog title 'platygastroidea' should be spelled as 'Platygastroidea' - the first letter 'P' should be capitalized] |
About
Writings related to insects, biodiversity and science in general Archives
July 2016
Categories
All
|





 RSS Feed
RSS Feed



